Found some more data:
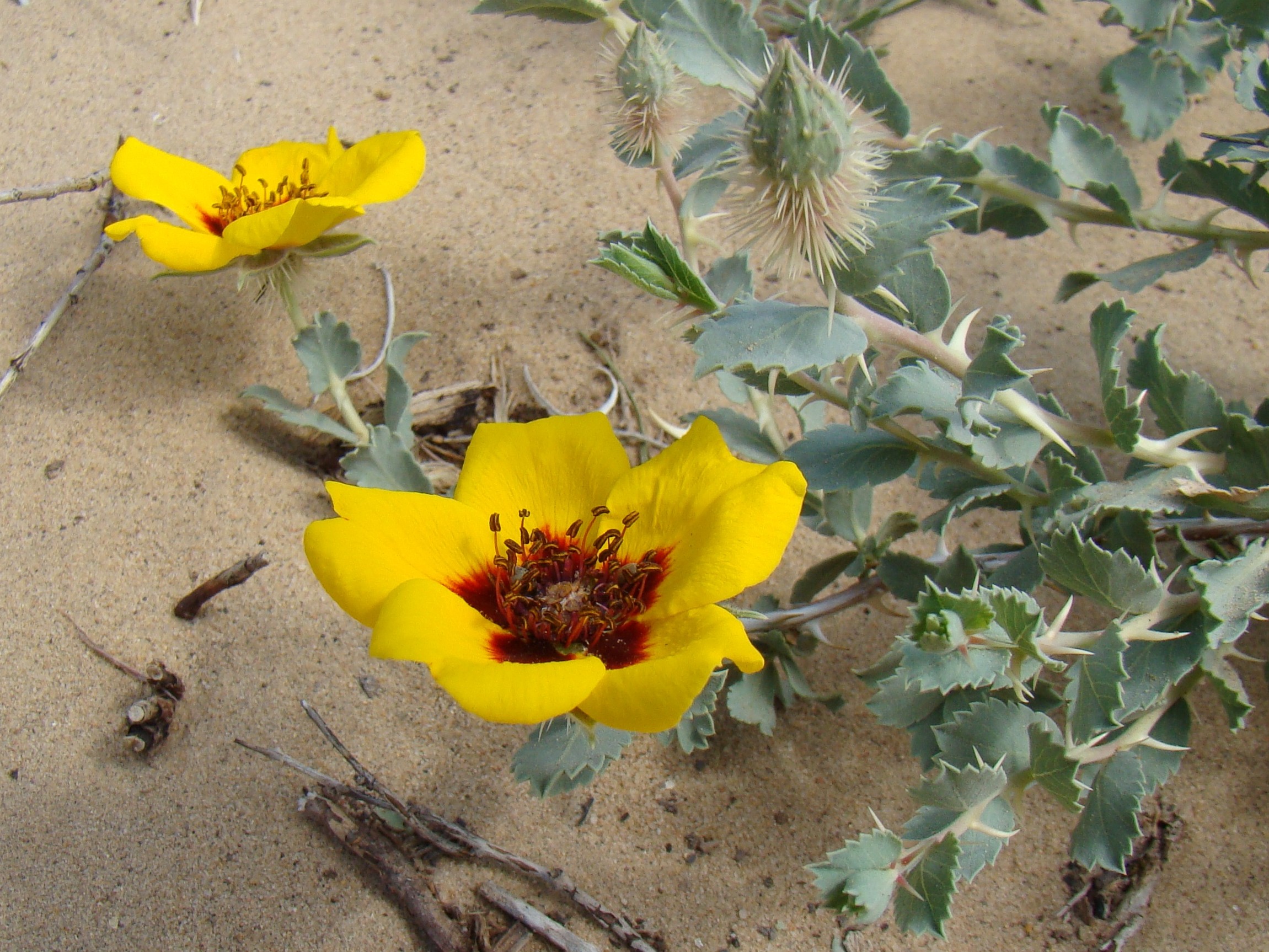
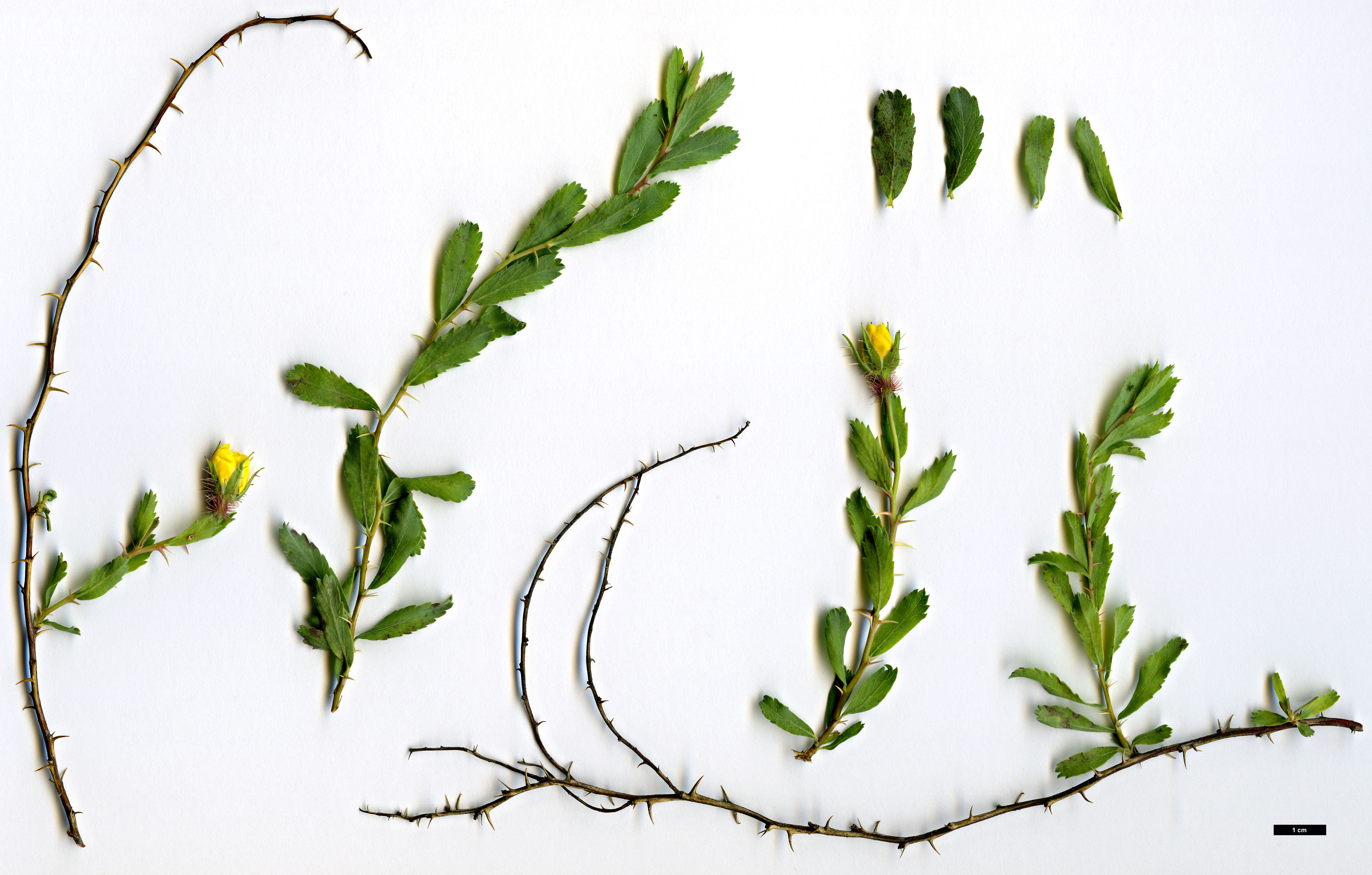
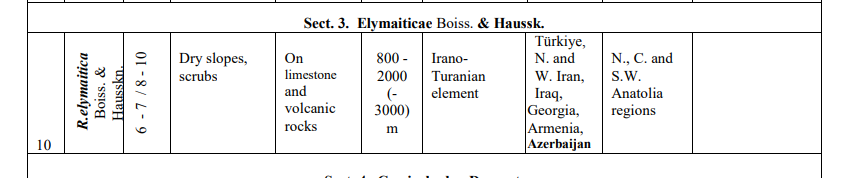
9. R. elymaitica , a bush up to 1 m high, with small pink (and rarely white) flowers, solitary or in clusters of 2-6 flowers; habitat: Hamadān, Kermānšāhān, Lorestān, Kohgiluya and Boir Aḥmad, Baḵtiāri, Isfahan, Fārs, Arāk, Qazvin, Tehran; also reported from Iraqi Kurdistan and eastern Anatolia (Zieliński, p. 19; Ḵātamsāz, pp. 54-56).
A page with Iranian plants: GOL – Encyclopaedia Iranica
And here you go  the mother load. I’ve done a deep dive in Google. I scientific study, some Persian original text and English translation. Apparently they also classify it under the Caninae Section. scientific studies and all.
the mother load. I’ve done a deep dive in Google. I scientific study, some Persian original text and English translation. Apparently they also classify it under the Caninae Section. scientific studies and all.
Page 94. There are 12 mentions of Rosa elymaitica. Apparently it is closely related to R. boissieri, R. orientalis and R. villosa.
Cluster analysis of all section Caninae showed an
extensive variance between seven Iranian species of the
section. There are two major cluster; populations of
R. boissieri and R. elymatica separated from the others in
the first cluster. The second cluster contained some sub- clusters that separated most of the populations of
R. orientalis and R. villosa. Population of R. canina, R. iberica, and R. pulverulenta distributed in both of
main groups and multivariate statistical data could not
segregate them. Polyploidy and cross pollination resulted
different hybrids between some species of section
Caninae as one of the extensive variation. The results of
ordination confirmed cluster analysis data (Fig. 6). Factor
analysis of morphological characters revealed that, the
first seven factors embraced about 60% of the total
variations, in which gland on adaxial leaflet, form of
sepal, hair on the adaxial leaflet, and pedicel form
could separate R. boissieri, R. elymatica, R. orientalis, and
R. villosa. Similarity between R. boissieri, R. elymatica, R. orientalis, and R. villosa was also considered by Flora
of Iran (Khatamsaz 1992). Morphometric data showed
some diversity in R. pulverulenta but in Flora of Iran
these species are placed near to each other which is not
supporting our data. Jokar et al. (2008) reported
R. pulverulenta is a hexaploid species. Moreover, there
are some reports on hybrids of R. pulverulenta and other
species such as R. pimpinellifolia (Khatamsaz 1992), and
R. boissieri. These data confirmed the diversity of
R. pulverulenta as our data showed. Morphological and
molecular studies between two subsections of Rosa
showed that, when different closely related dog-rose
species are present at the same growth site, the genetic
structure may show more differentiation between
localities than between taxa as it was shown in some
populations of R. pulverulenta in morphological
characters (De cock et al. 2008). From an evolutionary
point of view, the canina meiotic system has probably
developed fairly recently (Lim et al. 2005), supporting
the idea that, the dog-roses are rather young (Atienza et
al. 2005). Hexaploid base of R. Pulverulenta caused to
get equal molecular characters from their parents (not
maternal) and it could be reason of broad variation of
their populations.